Project 1 - Translatome changes in prion disease
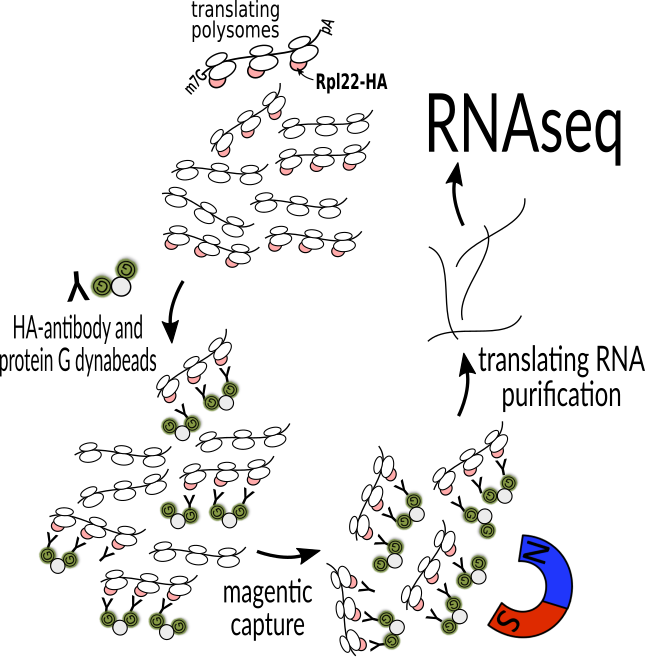
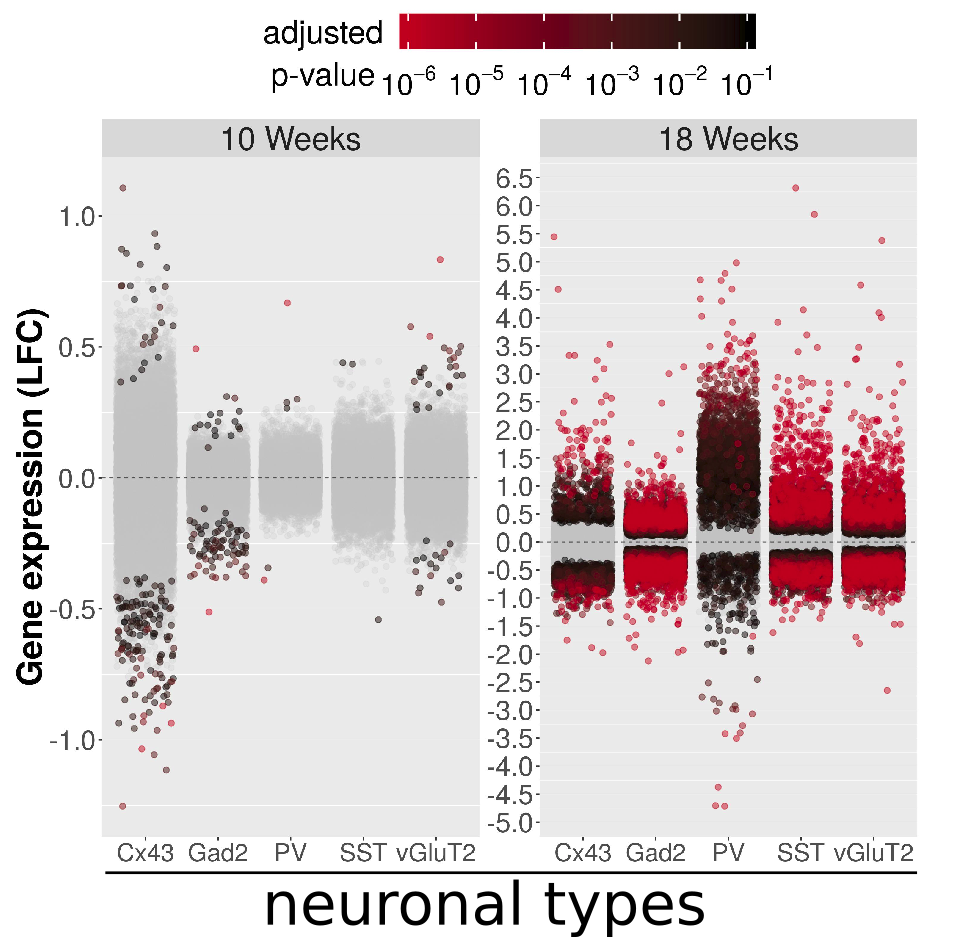
The RiboTag system enables us to study translation of mRNAs from specific types of cells in the mouse brain. This system is then applied to mouse models of neurodegenerative diseases. Applying RiboTag to a prion disease revealed that cells thought to be highly sensitive to disease do not reprogram their gene regulation system, while cells that are thought to resist disease do.
RiboTag in 3 genetic brain diseases
The initial success of project 1 has led us to study additional disease models, including Huntington’s disease (HD), Creutzfeldt-Jakob disease (CJD) and fatal familial insomnia (FFI). This project aims to study microglia, astrocytes, and four classes of neurons from the cerebellum, which is affected in the HD, FFI, and CJD models, and the rest of the brain, where different subregions are targeted for the different diseases.
Project 3 - A new tool: Tagger
We developed a new mouse line that enables us to study, from specific cell types, the full spectrum of gene regulation. With it we can study 1) the nucleus for changes to DNA methylation and histones, 2) pulse labeled RNAs, including lncRNAs and mRNAs, 3) miRNAs attached to Argonaute 2, and 4) the RiboTag.
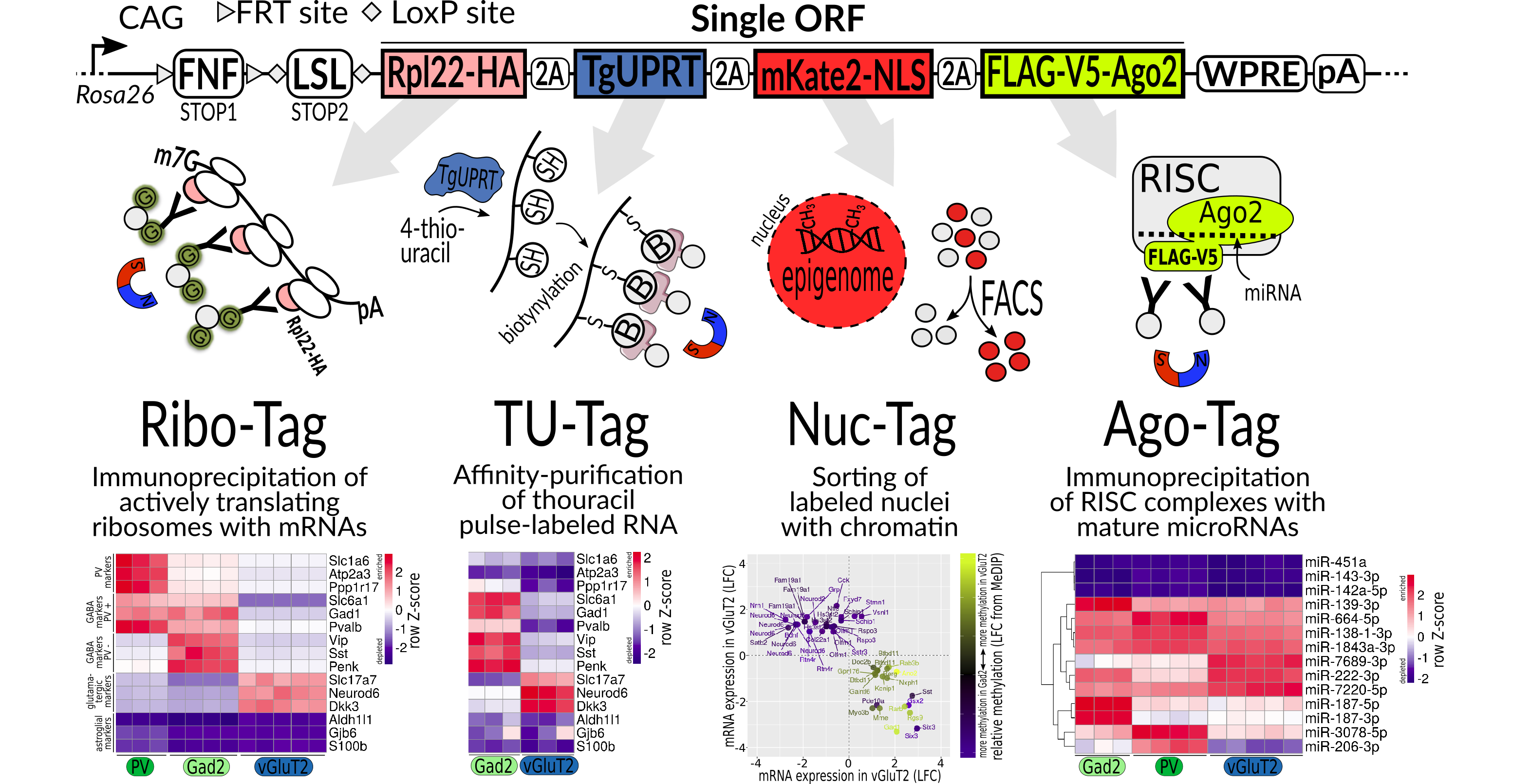
Project 4 - Applying Tagger to HD
HD is known to result in changes to gene regulation at many levels, including the chromatin, mRNA levels, translation, and miRNAs. We will use the Tagger system to study our HD model. We will study multiple ages and mutation sizes which correspond to differences in toxicity, and will focus on five different types of neurons in different brain regions.
Project 5 - Dynamics of gene regulation in vivo
The tagger system has us uniquely poised to study acute changes to gene regulation from specific cell types of complex tissues. A new computational method will make it possible to study dynamics or mRNAs that are captured on ribosomes. As models of acute changes to gene regulation, we will study the effects of the circadian rhythm and sleep deprivation with the Tagger system.